Abstract
Proliferation of wearable healthcare devices has created the need to deliver artificial intelligence applications for these resource-constrained devices to achieve faster, localized decision-making, by bringing computation closer to the data sources, for improved responsiveness and privacy. This contribution presents the results of an experimental evaluation of artificial neural network compression techniques, including quantization, structured pruning, and knowledge distillation, applied to multi-label classification of electrocardiogram (ECG) signals. The experiments were carried out on the PTB-XL dataset using three deep learning models, i.e. an LSTM-based recurrent neural network, a 1D convolutional neural network, and a 1D residual neural network. The results show how the compression methods impact model quality and highlight opportunities to reduce model size and accelerate inference, thereby enabling effective deployment on resource-constrained, edge devices.
1 Introduction
Resource-constrained environments refer to settings where limitations exist in computational power, memory, storage, or energy availability (Selvan
et al.,
2023). Examples include edge devices, such as wearable devices, IoT sensors, or embedded systems that require computational efficiency due to their restricted hardware capabilities or operational constraints, such as battery life or real-time processing needs. Constrained devices are common in mobile health applications, such as monitoring blood pressure and oxygen saturation, body temperature, breathing disorders, or heart conditions (Li
et al.,
2024).
At the same time, current artificial intelligence applications are often based on artificial neural networks (NNs) and require significant resources. Therefore, they may not be deployable in such environments. Model compression techniques attempt to address these issues and may enable intelligent applications to run on mobile devices with limited resources.
In this contribution, electrocardiogram (ECG) signal analysis was used to study the effects of model compression. Electrocardiography (Mirvis and Goldberger,
2001) is a fundamental non-invasive method of monitoring cardiac function. By placing electrodes on the patient’s skin, the ECG device captures the electrical signals of the heart that coordinate its behaviour during the cardiac cycle. ECG signals can be used, for example, to diagnose cardiovascular disease, detect sleep apnea, monitor heart rhythm, or to identify biometrics (Berkaya
et al.,
2018). Currently, there is an increase in the popularity of portable devices (such as smartwatches or chest bands) that are capable of capturing the ECG without access to medical facilities (Safdar
et al.,
2024), which allows for continuous monitoring of individuals’ health status.
Traditionally, physicians analyse ECG signals by evaluating wave patterns for abnormalities. Today, computational techniques, such as deep learning models, can be used to support diagnosis (Merdjanovska and Rashkovska,
2022). However, due to limited processing power and battery life, portable ECG devices often rely on data transfer and cloud-based infrastructures for data processing (Khan Mamun and Elfouly,
2023). Therefore, a key challenge is to adapt computationally demanding models to run on tiny devices (Khan Mamun and Elfouly,
2023). Here, the benefits include: (i) improved data privacy, (ii) real-time inference on the device, and (iii) reliable access to diagnostic tools in areas of low connectivity (Hohman
et al.,
2024). In this context, the potential of model compression techniques, namely quantization (Gholami
et al.,
2022), structured pruning (Cheng
et al.,
2024), and knowledge distillation (Hinton
et al.,
2015; Gou
et al.,
2021), can be explored to assess their effectiveness in reducing the computational demands of NN models.
Before proceeding, let us make several conceptual remarks. Although modern mobile devices are equipped with increasingly powerful hardware, this progress is counterbalanced by the growing size of NN models, which are trained on larger datasets and require more resources for inference. Moreover, the availability of devices’ computational power is also constrained, e.g. by the limitations in the battery capacity. Obviously, this limitation is not easy to overcome. Therefore, the focus of this contribution is to assess the effects of model compression techniques when applied to popular NN architectures, analysing the results that can be achieved. In doing so, it lays the foundation for further focused research. Separately, note that the ECG dataset serves as “sample data” that needs to be processed on resource-constrained devices, rather than being the focal point of this research.
Taking this into account, the key contributions of this work are as follows. (I) Three well-known deep learning model compression techniques were implemented, namely integer quantization, structured pruning, and response-based knowledge distillation suitable for the multi-label classification. (II) The methods were applied to three popular NN architectures commonly used for ECG analysis: a 1D CNN (convolutional neural network) (Lecun
et al.,
1998), an LSTM-based RNN (recurrent neural network) (Elman,
1990; Hochreiter and Schmidhuber,
1997), and a 1D ResNet (residual neural network) (He
et al.,
2016; Wang
et al.,
2017) to evaluate how the selected compression techniques improve resource utilization (measured in terms of inference speed and disk usage) and impact the quality of the results (measured in macro-averaged AUROC score). (III) The tests were performed using one of the largest open ECG datasets, PTB-XL (Wagner
et al.,
2020).
2 Related Work
Various NN compression techniques can be found in the literature (Li
et al.,
2023; Dantas
et al.,
2024). Their goal is to deliver a more compact (and more efficient) NN model that closely preserves the predictive quality of the original model. Quantization (Gholami
et al.,
2022) reduces the precision of numbers that represent the model parameters, typically from 32-bit floating-point numbers to 8-bit integers. Although it degrades model quality, it can lower the energy use and improve the inference speed (Hubara
et al.,
2016). Pruning (Cheng
et al.,
2024) removes “less important parts” of NNs, to decrease their size and computational load. Unstructured pruning discards individual weights scattered throughout a neural network and thus requires specialized hardware and software to take advantage of introduced sparsity (in terms of computation acceleration). On the other hand, structured pruning removes weights in a systematic way (i.e. it eliminates entire neurons, channels, filters, or layers), resulting in regular NNs, and thus is much more universal, because there is no need for dedicated hardware to obtain speed-up. Knowledge distillation (Hinton
et al.,
2015; Gou
et al.,
2021) transfers knowledge from a “teacher” model to a “student” model by training the student model to mimic the teacher’s output. The technique is typically used to create a student model smaller than its teacher.
For analysing ECG, deep learning models such as CNNs, ResNets, or RNNs are often used (Khan
et al.,
2023; Boulif
et al.,
2024; Safdar
et al.,
2024), but their computational demands can hinder deployment on portable or battery-powered devices. In the joint context of ECG analysis and NN compression, in (Lee
et al.,
2022), techniques such as pruning, quantization, and weight clustering were used to compress, e.g. ResNet, which detected arrhythmia based on ECG data, reducing the model’s size by a factor of 10000 with an accuracy loss of approximately 1%. The authors of (Chang
et al.,
2022) applied pruning and quantization to a CNN model for atrial fibrillation detection, achieving a 91-fold compression with an accuracy loss of roughly 1%. In Sepahvand and Abdali-Mohammadi (
2022), knowledge distillation was applied in an arrhythmia classification task, obtaining a model approximately 262 times smaller, while losing less than 1% of accuracy.
The extensive PTB-XL benchmark results are available in Strodthoff
et al. (
2020), where various NN architectures (including CNNs, ResNets, and RNNs) are compared in ECG tasks ranging from predicting diagnostic statements to determining age and gender. However, the benchmark does not consider the aspect of resource usage, which is adequate for the on-device ECG monitoring and analysis.
In this work, we conduct an experimental study of multiple compression strategies applied to the same ECG dataset. We systematically quantify how structured pruning, quantization, and knowledge distillation affect not only classification effectiveness but also model size and inference speed, showing the typical trade-offs involved in deploying deep ECG models on resource-constrained platforms.
3 Methodology
3.1 Dataset
To perform the experiments, the PTB-XL dataset (Wagner
et al.,
2020) was selected. It contained 21837 12-lead ECG 10-second recordings collected from 18885 patients. The recordings were available at sampling frequencies of 100 Hz and 500 Hz. They were supplemented with metadata, including identifiers, patient and measurement information, detailed diagnostic data, signal information, and recommended folds that split the data into training, validation, and test sets. Here, the diagnostic labels were divided into five categories: NORM (normal), MI (myocardial infarction), CD (conduction disturbances), HYP (hypertrophy), and STTC (ST-T changes). These labels could co-occur for a single recording.
For the experiments conducted in the presented research, the ECG time-series data were prepared for multi-label classification in the following way. Overall, standard approaches were followed (Berkaya
et al.,
2018). Recordings at the sampling frequency of 100 Hz were used. A fifth-order Butterworth high-pass filter (Butterworth,
1930) was used to remove low-frequency noise below 0.5 Hz. Power-line noise at 50 Hz was reduced using a bidirectional moving average filter. Each of the twelve leads was normalized separately. The training set was used to calculate the values needed for normalization, and then the remaining data were normalized using the computed statistics. The splits into training, validation, and test sets were performed using the recommended folds included in the dataset. Each recording was assigned a binary label vector
y of length 5, reflecting its membership in the diagnostic categories, as shown in (
1).
In accordance with the recommendation provided by the dataset authors (Wagner
et al.,
2020), the metric for assessing and comparing the models was the macro-averaged (the score was calculated for each label and then averaged) area under the receiver operating characteristic (AUROC) (Fawcett,
2006). This popular metric was selected because it measures the ability of a classifying model to discriminate between classes in a threshold-independent manner, eliminating the need to select a specific decision threshold (either globally or per model), on which many other metrics depend. AUROC has an intuitive probabilistic interpretation that its value corresponds to the probability that a classifier assigns a higher score to a randomly chosen positive instance than to a randomly chosen negative instance (Fernández
et al.,
2018).
3.2 Neural Networks
The following describes the NNs used in the experiments. The choice of the NN architectures was guided by their popularity in ECG classification. First, the RNN and the CNN architectures were created specifically for the purposes of this work, with their full, uncompressed versions intentionally designed to be compact (thus affecting the maximum compression rate). Next, the exact ResNet architecture was selected due to the superior benchmark results reported in Strodthoff
et al. (
2020). The chosen architecture types are generally common, supporting the usefulness of the gathered results. All NNs were implemented in PyTorch (version 2.2.2).
The RNN (Elman,
1990) was designed to capture the temporal dependencies inherent in ECG waveforms. The structure of the implemented RNN included an LSTM layer (Hochreiter and Schmidhuber,
1997) (with a hidden state dimension of 64), a layer normalization, a 1D max pooling layer (with stride and a kernel size of 125), a dropout layer (10%), a linear layer with 512 input features, and a sigmoid activation output. The 1D pooling layer divided the sequence of 1000 data points into eight segments (1.25 seconds each) and selected the maximum values. This reduced the data dimensionality and improved the results (by making the NN more robust to signal shifts). The model consisted of 22661 parameters. The Adam optimization algorithm was used, with an initial learning rate of
${10^{-3}}$ and a reduce-on-plateau scheduler that decreased the learning rate by half after every ten epochs without improvement, with a minimum learning rate of
${10^{-5}}$. The batch size was 64.
The CNN (Lecun
et al.,
1998) was created for hierarchical feature extraction from the input signals. The proposed CNN consisted of four convolutional blocks. Each block included a 1D convolution layer (kernel size of 3), a batch normalization layer, a ReLU activation function, and a 1D max pooling layer (kernel size of 3). The number of output channels, for each 1D convolution, was 32, 64, 96, and 32, respectively. The NN ended with a dropout layer (5%), a linear layer with 352 input features, and a sigmoid activation function. The model consisted of 37157 parameters. SGD with Nesterov momentum (
$\mu =0.995$) was used for training. The
$L2$ penalty coefficient,
λ, was experimentally set to 0.007. The learning rate was varied using a cosine annealing scheduler with a period of 30 epochs. The maximum learning rate was set at
${10^{-3}}$, while the minimum was set at
${10^{-6}}$. The learning rate was adjusted after each training epoch. The batch size was 64.
The ResNet (He
et al.,
2016) models were engineered to facilitate stable training in very deep models. The implementation of the 1D ResNet was based on the approach described in (Wang
et al.,
2017), with the following modifications. All shortcut connections were changed to 1D convolutional layers, with a filter length of 1 (He
et al.,
2016) (followed by a batch normalization) to enable pruning of all layers (Section
3.4). The output of the average pooling layer was passed to a linear layer with 128 input features, and the NN ended with a sigmoid activation function. The model consisted of 500869 parameters. The Adam optimizer was utilized. Cosine annealing was used to schedule the learning rate, with an annealing period of five epochs, a maximum learning rate of
${10^{-3}}$, and a minimum learning rate of
${10^{-6}}$. The batch size was 128.
To train the NNs for the multi-label classification, the cross-entropy loss (Goodfellow
et al.,
2016), shown in equation (
2), was used.
Here,
$n\in \mathbb{N}$ denotes the number of labels,
$y\in {\{0,1\}^{n}}$ represents the binary label vector from the dataset (Section
3.1), and
$p\in {(0;1)^{n}}$ is the vector of label occurrence scores returned by the model for the considered example.
3.3 Quantization
Let us now describe the applied compression techniques, starting with quantization. In this work, uniform post-training quantization (PTQ) (Gholami
et al.,
2022), expressed in (
3), was used. By default, static quantization was applied because it offers better inference time performance.
Here,
x represents the value to be quantized,
$q(x)$ denotes the quantized result,
α and
β define the range used for clipping values,
b specifies the number of bits of the quantized values,
z represents the zero point, and
r rounds its argument to the nearest integer value.
Inference, using quantized models, was run exclusively on a CPU. The quantization backend used a combination of FBGEMM and oneDNN libraries provided in PyTorch. Note that, in PyTorch, bias is not quantized.
The implementation utilized observers that collected information on floating-point tensors passing through the NN during calibration (Gholami
et al.,
2022; Wu
et al.,
2020), and derived quantization parameters (
α,
β,
z in (
3)) based on these statistics. For monitoring weights, PyTorch’s
PerChannelMinMaxObserver was used. It identified the minimum and maximum values for each channel, along the specified axis, to derive quantization parameters. The weights followed a symmetric quantization scheme (Gholami
et al.,
2022) with per-channel granularity (Wu
et al.,
2020). They were represented as 8-bit signed integers, with range
$[-127;127]$ (sacrificing one value for symmetry).
The activation observer was PyTorch’s
HistogramObserver, which minimized the error between the floating-point and the quantized data distributions. Layer activations used the asymmetric quantization scheme (Gholami
et al.,
2022) with per-tensor granularity (Wu
et al.,
2020). Their quantized form was represented using 8-bit unsigned integers, although 7 bits were used (range 0 to 127) to avoid saturation (Li and Alvarez,
2021) caused by the assembler instruction
VPMADDUBSW in the FBGEMM library implementation (AVX-512 VNNI instructions were not available on the CPU used). The exception was the output sigmoid activation function, which used the full 8-bit range.
The LSTM layer was handled differently. PyTorch’s
MinMaxObserver was used to determine the quantization parameters for the weights. It found the quantization parameters collectively for all weights at once (using static quantization). Thus, it was per-tensor granularity, where the quantization parameters were shared for all weights. The remaining weight quantization configuration was the same as before. Here, activations were dynamically quantized (Gholami
et al.,
2022) during inference, so the quantization parameters were based on the values observed at that moment. The remaining activation quantization configuration was unchanged.
Additionally, operator fusion was used to combine the convolutional layer, the batch normalization layer, and the ReLU activation function (if it occurred in the sequence) into a single layer. Finally, 128 randomly selected examples from the validation dataset were used to calibrate all models.
3.4 Pruning
A structured variant of pruning (Cheng
et al.,
2024) was chosen, because it does not require specific hardware to observe the acceleration benefits. It was implemented for the NNs proposed in Section
3.2.
The LSTM (Hochreiter and Schmidhuber,
1997) layer consists of the input-hidden weights
${W_{xi}}$,
${W_{xf}}$,
${W_{xc}}$,
${W_{xo}}$ $\in {\mathbb{R}^{r\times d}}$ and the hidden-hidden weights
${W_{hi}}$,
${W_{hf}}$,
${W_{hc}}$,
${W_{ho}}$ $\in {\mathbb{R}^{r\times r}}$. Where
$d\in \mathbb{N}$ is the number of input features and
$r\in \mathbb{N}$ is the number of features in the hidden state. At time step
$t\in \mathbb{N}$, the weights are used to compute the input gate in (
4), the forget gate in (
5), the cell gate in (
6), and the output gate in (
7). Here,
${x^{(t)}}\in {\mathbb{R}^{d}}$ is the input vector at time
t, and
${h^{(t-1)}}\in {\mathbb{R}^{r}}$ is the hidden state at time
$t-1$. For brevity, the bias terms (
b) are omitted from the considerations because they follow the pruning pattern derived from the weights in the solution.
Therefore, the weights related to the
s-th neuron in the input, forget, cell, and output gates can be expressed as
${W_{s}}\in {\mathbb{R}^{4\times (d+r)}}$ in (
8), where
$s\in \{1,\dots ,r\}$.
For each
${W_{s}}$, either the
$L1$ or the
$L2$ norm was calculated. The neurons with the lowest values were pruned (thus, the weights and the biases associated with these neurons were removed).
An approach similar to (Li
et al.,
2017) was applied to prune the convolutional layers. Consider a 1D convolutional layer, where weights are stored in a 3D tensor of size
$O\times I\times K$. Here,
O is the number of output channels,
I is the number of input channels, and
K is the length of the filter (the bias terms are again omitted for brevity). Layer output channels are formed independently, so let us consider a single output channel, a matrix
${W_{s}}$ of size
$I\times K$, where
$s\in \{1,\dots ,O\}$. Now, suppose that
$u\in \{0,\dots ,I-1\}$ input channels become unavailable. Then, these missing channels carry no information. Hence, effectively, the matrix
${W_{s}}$ becomes
${W^{\prime }_{s}}$ of size
$(I-u)\times K$, without the rows corresponding to the unavailable channels. During pruning,
${W^{\prime }_{s}}$ were considered, thus avoiding missing input channels from earlier layers (Li
et al.,
2017). Similarly to the LSTM layer, for
${W^{\prime }_{s}}$ values, the norm
$L1$ or
$L2$ was calculated. Then, the layer’s output channels with the smallest values were pruned. In the ResNet, its shortcut connections followed the pruning pattern of the last convolutional layer in their residual blocks.
The pruning procedure started from the input layer and proceeded sequentially until the output layer. For the RNN model, pruning began with the LSTM layer. Based on the norm values, calculated for
${W_{s}}$ (
8), neurons were removed, and the pruning schema was passed to the rest of the NN. For the CNN and the ResNet, in a convolutional layer, after receiving information about the unavailable input channels, the selected norm of
${W^{\prime }_{s}}$ was computed, and the output channels, with the lowest norm values, were removed and the pruning schema was passed down deeper into the NN. The remaining layers, such as the batch normalization layers, the linear layers, etc., reacted to the missing channels by pruning the appropriate weights and biases, in effect, shrinking as well.
Ideally, the model reduction achieved with structured pruning should lower the computational load, e.g. measured in simple hardware-agnostic proxy metrics such as FLOPs (floating-point operations), making the inference faster. However, in practice, structured pruning may lead to a suboptimal configuration of NN structures (Dong
et al.,
2021; Liberis
et al.,
2021). This suboptimality manifests itself in not utilizing the full potential of the underlying hardware and the library implementations, e.g. highly optimized vectorized kernels, effectively leading to notable slowdowns.
During the experiments, pruning was performed in one or more rounds to achieve the same degree of structural reduction measured in the fraction of structures left in the final pruned model. The fraction of available structures pruned in a single layer in one round was expressed as $p=1-{f^{1/r}}$, where $r\in \mathbb{N}$ denoted the number of pruning rounds and $f\in (0;1)$ was the target fraction of structures left after all rounds (note the difference between weights and structures). In the solution, f was set to 75%, 50%, and 12.5%; r was 1, 5, or 10.
After each pruning round, the models were fine-tuned, using generally the same approach as for training the full (uncompressed) NNs, or using knowledge distillation (Section
3.5). However, for the CNN model, the maximum learning rate was reduced to
$5\cdot {10^{-4}}$ and the annealing period was changed to 20 epochs. The RNN and the ResNet configurations were unchanged.
3.5 Knowledge Distillation
Overall, response-based knowledge distillation (Gou
et al.,
2021) was used, offering a simple and general approach. However, the typical scheme introduced in (Hinton
et al.,
2015) suits multi-class, not multi-label classification. Thus, the authors of (Yang
et al.,
2023) proposed applying response-based knowledge distillation with one teacher and one student to multi-label classification, by decomposing the multi-label task into independent binary classification tasks. In these tasks, two two-element probability distributions (of the student and the teacher) are created by taking the probability of label occurrence and its complement and, finally, expressing the loss function as the divergence (
$\mathcal{D}$) between these distributions. This loss function,
${\mathcal{L}_{MLD}}$ (Yang
et al.,
2023), is presented in (
9), where the superscript
t denotes the teacher’s output and
s, the student’s output. See (
2) for the remaining notation.
Ultimately, the loss function followed expression (
10). In
${\mathcal{L}_{CE}}$ (
2),
p was set to be the student’s output,
${p^{s}}$. As divergence,
$\mathcal{D}$, in (
9), Kullback-Leibler divergence (Kullback and Leibler,
1951) was used. For all models, the best coefficient
α was empirically determined to be 0.4.
During the experiments, the teacher model was a full floating-point model trained without knowledge distillation with the same base architecture as the student. The student was either a new instance of an uncompressed model to see if the approach can improve the base models’ results, or a pruned (smaller) model to observe the impact during the fine-tuning stage.
3.6 Additional Details of the Experimental Setup
For memory allocation, instead of malloc, tcmalloc was used for the RNN, and jemalloc for the CNN and the ResNet models, as they offered slightly faster performance.
The average inference time was calculated for the models running on the CPU (Section
4). Although it may not be correlated with the execution on other hardware, clock time was selected in order to show the actual latency that captures various factors, such as memory access or kernel launch overheads. The number of threads available to PyTorch was limited to 1 to mitigate the influence of parallel execution, which was beyond the scope of this research.
Before the time measurements, the models were warmed up on 100 examples. The batch size was set to 1. Inference time was measured for 10000 examples for the CNN, and 2000 examples for the RNN and the ResNet, and the results were averaged. In PyTorch, the floating-point version of the RNN model can use an efficient implementation for the LSTM layer from the oneDNN library (formerly known as MKL-DNN and DNNL). Therefore, the floating-point RNN inference time was measured twice, with and without access to the library.
The models’ size was measured by storing PyTorch’s state_dict on the disk and then obtaining its size. Note that, in this representation, additional data are stored in the created file, such as information about tensors representing weights or quantization parameters. Therefore, the reported size was worse than the ideal theoretical reduction (i.e. a 4-fold reduction in the case of quantizing the model from 32-bit floating-point to 8-bit integer representation).
4 Experiments and Results
Across this section, the following notation is used. M${_{\text{F}}}$ is a floating-point model; M${_{\text{Q}}}$ is the quantized M${_{\text{F}}}$; M${_{\text{FKD}}}$ is a floating-point model trained with knowledge distillation, where the full M${_{\text{F}}}$ was the teacher; M${_{\text{QKD}}}$ is the quantized M${_{\text{FKD}}}$. The notation $r/Ln$ means that the model results from pruning in the total of r rounds using the $Ln$ norm ($L1$ or $L2$). The numerical results are reported as the mean value, with the standard deviation, presented as 0.abcd(xyz), which should be understood as 0.abcd ± 0.0xyz, e.g. 0.9186(013) stands for 0.9186 ± 0.0013. The term “parameters left” refers to the fraction of the weights and the biases left in the model after pruning.
The experimental process was as follows. First, a basic model, M
${_{\text{F}}}$, was trained. Next, a new model, M
${_{\text{FKD}}}$, was trained using M
${_{\text{F}}}$ as the teacher in the knowledge distillation schema. After that, M
${_{\text{F}}}$, or M
${_{\text{FKD}}}$, was pruned in
r rounds using the
$L1$ or the
$L2$ norm, with each round followed by fine-tuning (with knowledge distillation for M
${_{\text{FKD}}}$). Throughout the process, models reached their target stage; for not pruned models, that was the end of the initial training, while for pruned models, it was the point where the desired target fraction of structures left,
f (Section
3.4), was achieved. After reaching the target stage, each resulting model was quantized, producing M
${_{\text{Q}}}$ or M
${_{\text{QKD}}}$, and tested.
The experiments were run five times for each architecture. Tables
1,
2, and
3 present the mean (from the five runs) macro-averaged AUROC in each target stage, achieved by the full model (100% of parameters) or by the best pruning approach. The highest score in each row is written in bold, and the highest score in the table is additionally underscored. Figures
1,
3,
5 with inference time and Figs.
2,
4,
6 with the models’ size contain the averaged measurements from the five runs, along with the error bars representing the standard deviation. These measurements were performed for each intermediate model produced during the experiments.
To accelerate development and collect preliminary results, the experiments were conducted on the Intel i9-12900H CPU and the NVIDIA GeForce RTX 3080Ti Laptop GPU rather than on an ECG device (Section
5.5). For consistency, the CPU scaling governor was set to performance mode and Intel Turbo Boost was enabled.
4.1 Recurrent Neural Network
Table 1
RNN AUROC scores with pruning techniques.
| Model |
Parameters left |
| 100% |
61.45% |
31.94% |
4.61% |
| M${_{\text{F}}}$
|
0.9186(013) |
0.9199(009) |
0.9186(005) |
0.8755(091) |
|
1/$L1$
|
10/$L2$
|
10/$L1$
|
| M${_{\text{Q}}}$
|
0.9156(008) |
0.9155(010) |
0.9136(010) |
0.8697(101) |
|
1/$L1$
|
10/$L2$
|
10/$L1$
|
| M${_{\text{FKD}}}$
|
0.9207(005) |
0.9208(011) |
0.9195(021) |
0.8809(058) |
|
10/$L2$
|
5/$L2$
|
5/$L1$
|
| M${_{\text{QKD}}}$
|
0.9174(008) |
0.9178(008) |
0.9167(014) |
0.8724(070) |
|
10/$L2$
|
10/$L2$
|
1/$L2$
|
What follows is a summary of the results obtained by the RNN during the experiments. First, we analyse the AUROC results in Table
1. Models M
${_{\text{F}}}$, M
${_{\text{FKD}}}$, and M
${_{\text{QKD}}}$ achieved their best scores at 61.45% of the original parameters left, demonstrating that the applied pruning method (both with and without knowledge distillation) has the potential of improving the quality of the base model. This behaviour can be attributed to the regularizing effect of pruning (Hohman
et al.,
2024). However, more aggressive parameter reduction that shrank the models to 31.94% of parameters resulted in degraded quality, and further reduction to 4.61% led to a poor fit of the models. Therefore, practitioners must carefully consider the trade-off between the model size and the predictive quality while applying pruning.
The models fine-tuned with knowledge distillation (M${_{\text{FKD}}}$, M${_{\text{QKD}}}$) outperformed their respective counterparts fine-tuned without knowledge distillation (M${_{\text{F}}}$, M${_{\text{Q}}}$) in terms of predictive quality across all pruning levels. Interestingly, for the models M${_{\text{FKD}}}$ and M${_{\text{QKD}}}$ at 31.94% and 4.61% different pruning approaches resulted in highest scores (i.e. 5/$L2$ versus 10/$L2$, and 5/$L1$ versus 1/$L2$). This observation indicate that the best quantized model does not necessarily derive from the highest-scoring floating-point model.
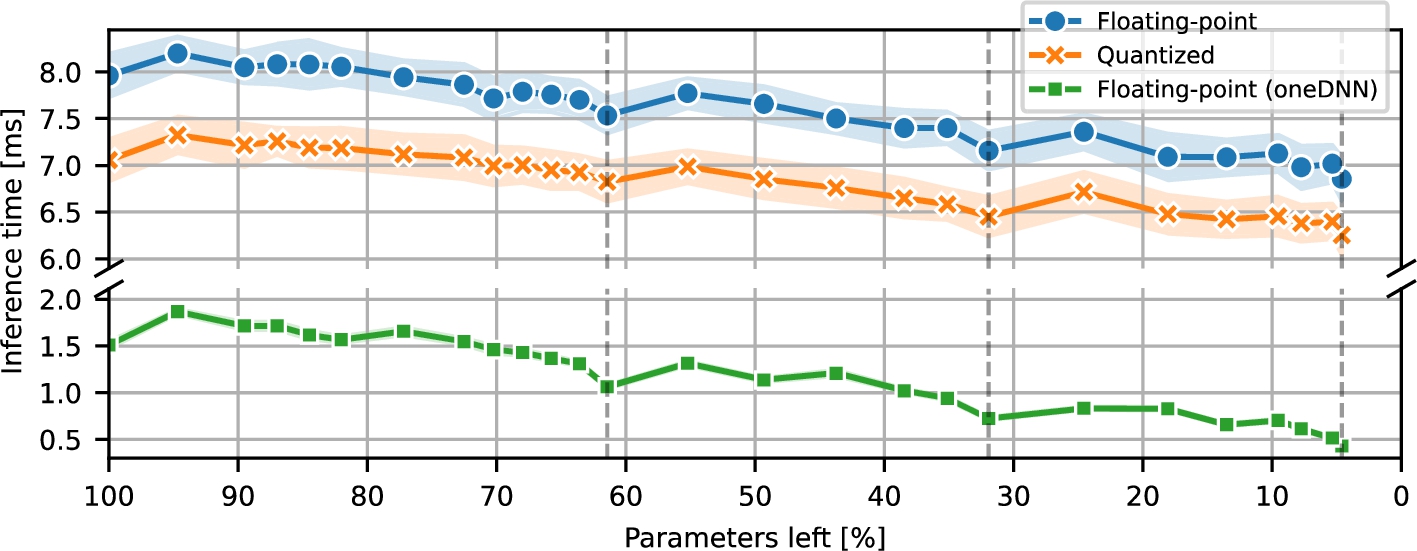
Fig. 1
RNN inference time during pruning.
Next, Fig.
1 presents the average inference time of the both floating-point (M
${_{\text{F}}}$, M
${_{\text{FKD}}}$) and the quantized (M
${_{\text{Q}}}$, M
${_{\text{QKD}}}$) models during pruning. The time measurements of the floating-point models were performed with and without access to the oneDNN library (Section
3.6). As expected, the floating-point models without access to the oneDNN implementation were slower than their quantized counterparts. Moreover, the floating-point models using the oneDNN library were significantly faster than the other two (those without access to the library and quantized ones), thus showing that quantization may not always be universally advantageous in terms of performance. However, it has to be stressed that this conclusion is likely to be platform-specific. Additionally, since during pruning, the absolute differences of latency improvements for floating-point models were similar to each other and more significant than the ones observed in the quantized models, the biggest relative improvements were observed for the floating-point models with access to the oneDNN library.
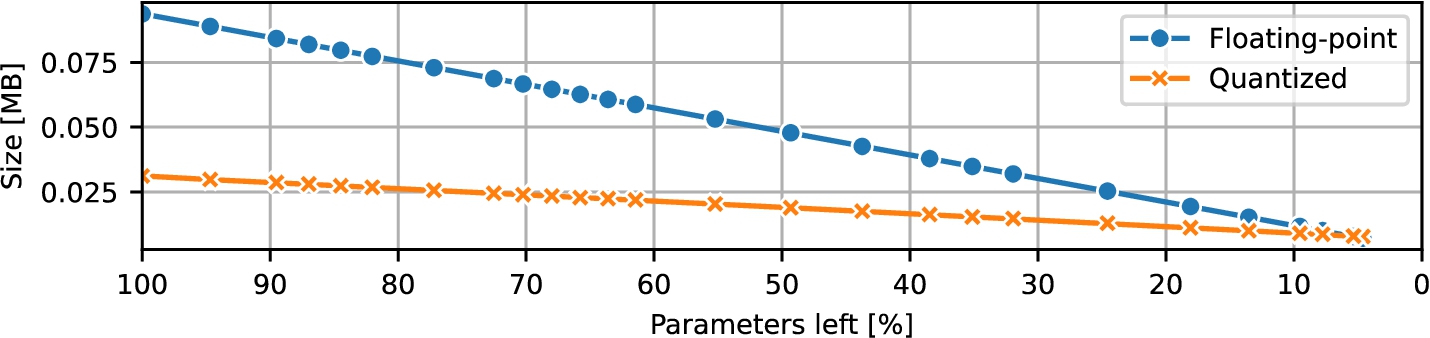
Fig. 2
RNN size during pruning.
During pruning, the size of the models decreased linearly; the same was true also for the CNN and the ResNet. The floating-point RNN models shrank from 93.677 KB to 7.149 KB, while the quantized models decreased from 31.194 KB to 7.770 KB. Therefore, the storage advantage of using quantization gradually diminished with size, to a certain point where the quantized NN actually consumed more disk space, due to additional overhead data stored on the disk (Section
3.6). Figure
2 visualizes these size measurements throughout pruning.
4.2 Convolutional Neural Network
Table 2
CNN AUROC scores with pruning techniques.
| Model |
Parameters left |
| 100% |
57.95% |
27.27% |
2.56% |
| M${_{\text{F}}}$
|
0.9311(006) |
0.9309(003) |
0.9302(003) |
0.8984(031) |
|
10/$L2$
|
10/$L2$
|
5/$L1$
|
| M${_{\text{Q}}}$
|
0.9253(028) |
0.9227(056) |
0.9216(036) |
0.8865(081) |
|
1/$L1$
|
5/$L1$
|
10/$L1$
|
| M${_{\text{FKD}}}$
|
0.9320(007) |
0.9311(005) |
0.9299(011) |
0.9032(037) |
|
10/$L2$
|
5/$L2$
|
10/$L2$
|
| M${_{\text{QKD}}}$
|
0.9268(019) |
0.9252(036) |
0.9242(016) |
0.8808(187) |
|
5/$L2$
|
1/$L2$
|
10/$L1$
|
Next, we discuss the results obtained by the subsequent proposed model, CNN. The data in Table
2 summarize the AUROC scores obtained by the CNN at different target stages. In contrast to the patterns observed for the RNN (Section
4.1) and ResNet (Section
4.3), the CNN models suffered a noticeable drop in AUROC scores once parameters were removed. The smallest degradation appeared in the models trained with knowledge distillation, suggesting that it helped the pruned networks to retain part of the teacher’s representational power. However, the benefit was not uniform. At 27.27% parameters left, the model fine-tuned with knowledge distillation, M
${_{\text{FKD}}}$, was slightly wore than its non-distilled counterpart, M
${_{\text{F}}}$. Furthermore, at a high sparsity level of 2.56% of parameters left, M
${_{\text{Q}}}$ was a bit better than M
${_{\text{QKD}}}$. Another aspect to note in the results is that different combinations of pruning rounds and norms led to the best-performing floating-point and quantized models at the same pruning level. For instance, at 57.95% of parameters left, the best results for the floating-point model, M
${_{\text{F}}}$, were achieved with 10/
$L2$ pruning method, but the best results for the quantized model, M
${_{\text{Q}}}$, were achieved after quantizing the floating-point model obtained with 1/
$L1$ pruning method. Therefore, one cannot always assume that among multiple floating-point models, the one with the best results will still be the best after quantization.
Although relatively small and compact, the CNN architecture used in the reported experiments turned out to be well-performing on the given (ECG) dataset. In fact, pruning the CNN so that only 27.27% of its original parameters remained, it still outperformed the best RNN results. Even more encouraging is that the base M
${_{\text{FKD}}}$ CNN reached the AUROC score that surpassed the best single model scores reported by (Strodthoff
et al.,
2020) for the very same task. These findings suggest that a carefully designed, lightweight CNN can match (or even beat) larger and more complex models on ECG classification while using a fraction of the parameters, highlighting the importance of tailored architecture design.
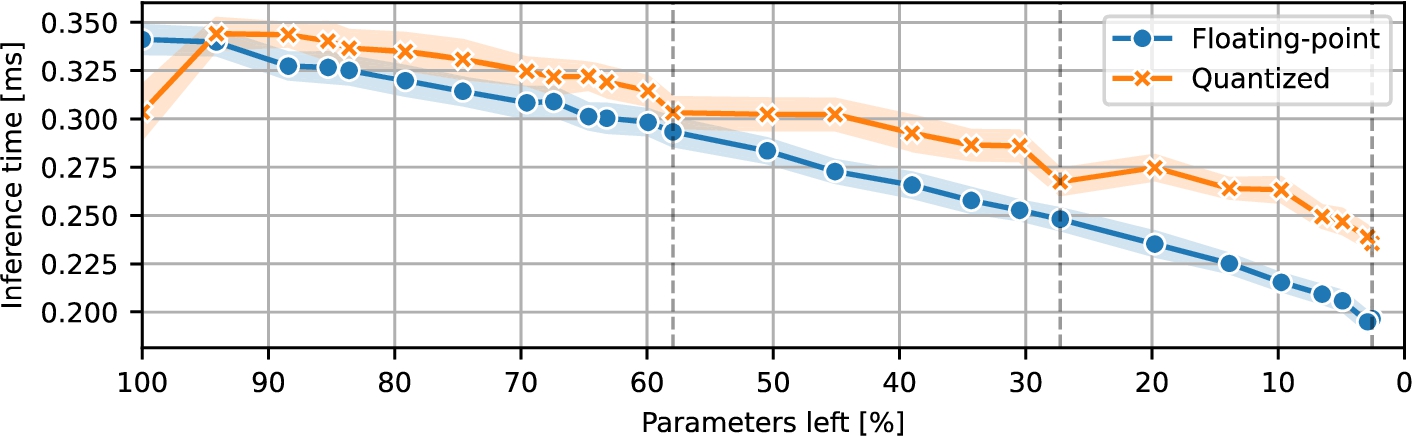
Fig. 3
CNN inference time during pruning.
Next, Fig.
3 illustrates the inference latency of the CNN variants in the experiments. Initially, the quantized models outperformed their floating-point counterparts, achieving lower inference times on the tested hardware. However, during pruning, the trend shows that the models performed worse after quantization, mainly due to the degraded performance of the quantized max pooling operation with less than 32 channels, causing quantized models to exhibit higher latency than the floating-point versions despite their reduced arithmetic precision.
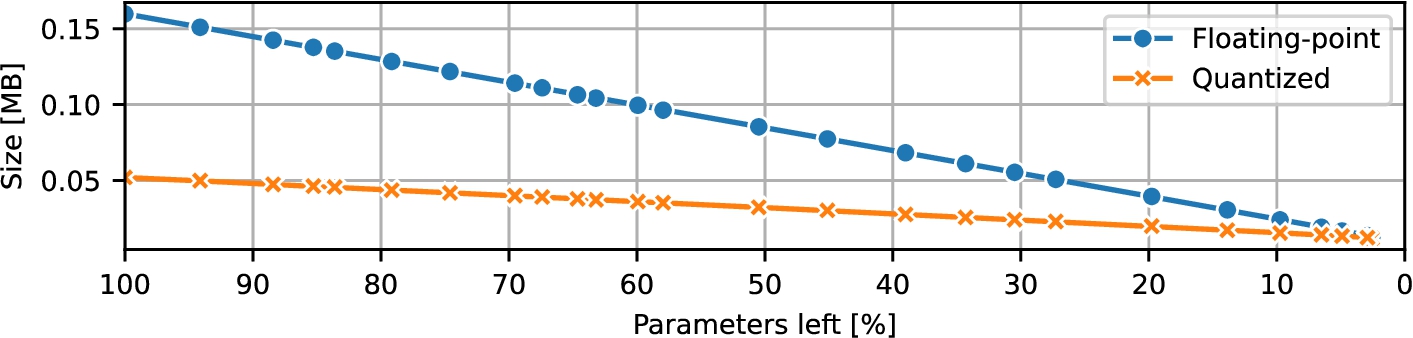
Fig. 4
CNN size during pruning.
The models’ on-disk size measurements are presented in Fig.
4. The original floating-point networks occupied 159.734 KB; the smallest models required only 13.174 KB of storage. The quantized networks started at 51.962 KB and were pruned down to 11.898 KB. Thus, even though the pruned quantized models incurred a latency penalty, they consistently maintained a smaller on-disk size compared to their floating-point equivalents.
4.3 Residual Neural Network
Table 3
ResNet AUROC scores with pruning techniques.
| Model |
Parameters left |
| 100% |
56.60% |
25.47% |
1.77% |
| M${_{\text{F}}}$
|
0.9282(006) |
0.9283(010) |
0.9274(008) |
0.9228(011) |
|
1/$L1$
|
1/$L1$
|
10/$L2$
|
| M${_{\text{Q}}}$
|
0.9093(101) |
0.9107(065) |
0.9069(101) |
0.8936(103) |
|
1/$L1$
|
1/$L1$
|
1/$L2$
|
| M${_{\text{FKD}}}$
|
0.9292(004) |
0.9297(010) |
0.9294(009) |
0.9257(009) |
|
10/$L1$
|
10/$L2$
|
10/$L2$
|
| M${_{\text{QKD}}}$
|
0.9193(047) |
0.9183(039) |
0.9122(082) |
0.9053(111) |
|
5/$L1$
|
5/$L1$
|
10/$L2$
|
The following section summarizes the AUROC results obtained by the proposed ResNet-based architecture (Table
3). Several insights emerge when we compare unpruned baselines, pruned variants, and models trained with knowledge distillation. It is also worth mentioning that this (baseline) model contained significantly more parameters than the other two analysed NNs.
First, pruning positively impacted some of the scores. The removal of about half of the parameters yielded slight AUROC improvements for models M
${_{\text{F}}}$, M
${_{\text{Q}}}$, M
${_{\text{FKD}}}$. Similar to the RNN, this increase can be attributed to the regularizing effect of pruning, which mitigates overfitting and encourages the network to focus on the most informative feature maps. In contrast, M
${_{\text{QKD}}}$ experienced a slight drop in AUROC following a comparable pruning. Interestingly, it was observed that M
${_{\text{F}}}$ pruned in a single round to 56.60% and 25.47% of parameters left performed better than the same model pruned (and fine-tuned) in 5 or 10 rounds, which led to over-fitting; this was not observed for M
${_{\text{FKD}}}$. However, M
${_{\text{F}}}$ and M
${_{\text{FKD}}}$ shared a mutual behaviour for much more disruptive pruning, to 1.77% parameters left, where a gradual approach with more rounds offered superior results compared to faster pruning in fewer rounds. Moreover, across the experiments, the model variants obtained using knowledge distillation (M
${_{\text{FKD}}}$ and M
${_{\text{QKD}}}$) consistently surpassed their non-distilled counterparts. This behaviour demonstrated that transferring label information during training not only enhances generalization but also protects the models from the adverse effects of parameter removal. Finally, it is worth noting that the unpruned baseline M
${_{\text{F}}}$ achieved the AUROC score of 0.9282, matching the performance reported in (Strodthoff
et al.,
2020) for the same task using an analogous model that scored 0.930. This parity can be treated as a confirmation of the validity of the proposed implementation.
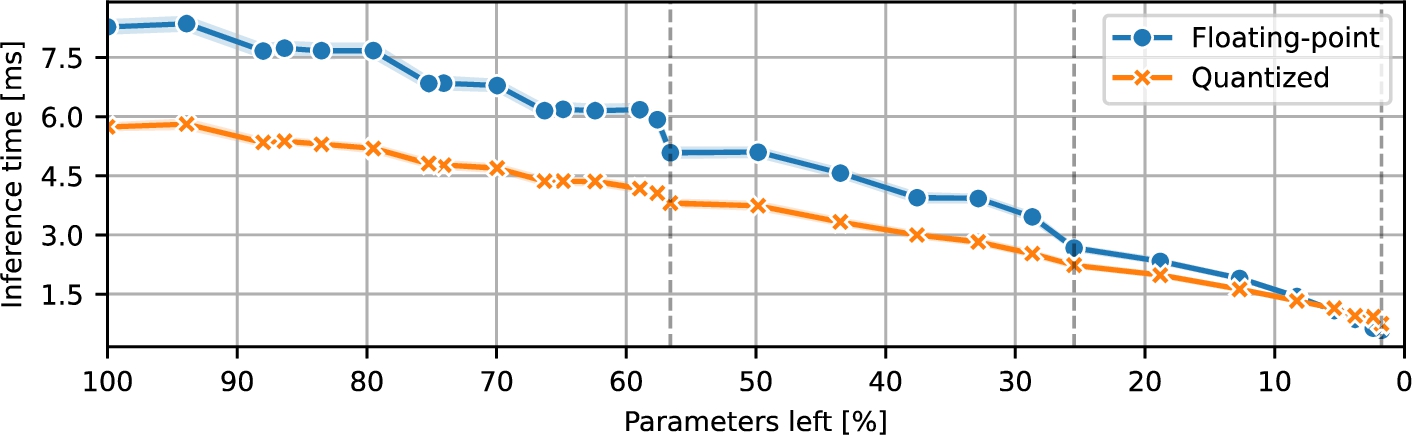
Fig. 5
ResNet inference time during pruning.
Next, Fig.
5 shows how inference latency for ResNet models changed during the experiments. For the vast majority of pruning levels, the quantized models were noticeably faster than their 32-bit floating-point counterparts; the reduced arithmetic complexity translated directly into lower inference times. However, near the maximum level of pruning, when the remaining parameter count (and thus model size) of the ResNet is roughly on par with that of the analysed baseline CNN (both architectures were convolution-oriented), the observed trend flipped. In this ultra-pruned regime, the quantized ResNet actually incurred slightly higher latency than the floating-point version, a behaviour that mirrored the pattern observed for the CNN in Fig.
3.
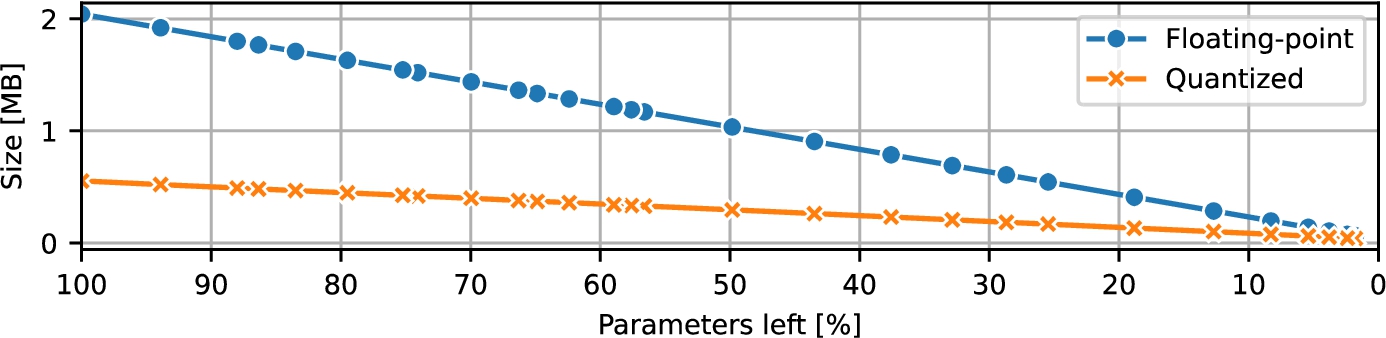
Fig. 6
ResNet size during pruning.
The initial floating-point models occupied 2.042902 MB at their largest and were pruned down to just 0.065302 MB in their most compact form. As anticipated, applying quantization reduced these footprints even further, and the quantized models consistently outpaced their floating-point counterparts in storage efficiency. Their size ranged from 0.553758 MB (100% of parameters) to 0.041374 MB (1.77% of parameters left). The trends are visualized in Fig.
6.
5 Discussion
5.1 Neural Networks
The uncompressed, full-precision versions of each proposed NN achieved classification performance (measured by macro-averaged AUROC) that closely matched or slightly exceeded the benchmark results reported in Strodthoff
et al. (
2020). Moreover, these types of NNs represent different computation characteristics that range from the sequential processing of RNNs to the shortcut-connected layers of ResNets. Therefore, they were deemed good candidates for the subsequent experimentation with the compression techniques, which were the focal point of this work.
5.2 Quantization
In the conducted experiments, in almost all tested cases but one, quantization reduced the models’ storage footprint owing to the use of lower precision numbers to represent the weights, i.e. 8-bit integers instead of 32-bit floating-point parameters. In theory, the use of this kind of quantization should result in a four-fold reduction in the on-disk size storage requirements, yet in practice, we observed smaller (worse) savings. Moreover, for the heavily pruned RNN, the quantization was detrimental size-wise, i.e. it resulted in a bigger footprint than the full-precision model. Both the discrepancy between the actual and four-fold reduction and the RNN exception were traced back to the implementation details of PyTorch and the method used to measure the disk usage (Section
3.6).
Despite making the models smaller and using a representation that theoretically offers faster computation with integer arithmetic, the effect of quantization on the inference speed on the tested CPU was variable, did not always result in faster inference, and heavily depended on the model architecture, the NN size, and the libraries used. Quantized RNN models were faster than their full-precision counterparts. However, using the oneDNN library for the floating-point models proved to be much faster and outperformed even the integer variants. Next, quantized CNN models experienced a slowdown at the beginning of the pruning. Although the inference speed kept improving, they remained slower than the full-precision models. In effect, during pruning, the floating-point CNNs were superior to the quantized models in terms of inference speed. The ResNet architecture, which was the largest network in the experiments, benefited the most from the quantization. However, once it reached a size comparable to that of the used CNNs, a similar slowdown occurred that resulted in the quantized models being slower than their full-precision counterparts. These results underscore that the impact of quantization on speed is highly context-dependent. However, it is important to note that the results could also differ on other hardware, e.g. without a dedicated floating-point unit, or in other runtime environments, e.g. due to the specifics of quantized kernel implementations.
As expected, replacing floating-point 32-bit weights with quantized integers degraded the predictive quality of the tested models. However, the exact magnitude of impact on the AUROC score depended on the NN architecture and the NN size. The largest considered model, ResNet, suffered the biggest negative impact on the relative scores between non-quantized and quantized models. At the same time, the most negligible negative effect was observed for the RNN architecture, which had the smallest number of parameters. In the case of the CNN architecture, the penalty resulting from quantization was between the other two architectures. Nonetheless, in all cases, the loss in predictive quality was gradual rather than catastrophic. Potentially, more sophisticated calibration or quantization-aware training could help recover part of the gap.
5.3 Pruning
Structured pruning was naturally beneficial for reducing the models’ disk usage because, in the proposed implementation, the pruned structures were actually removed from the network and not merely masked. As expected, the decrease followed a linear trend as a function of the number of parameters left in the model.
Beyond storage savings, structured pruning yielded improvements in inference latency across all architectures, although the magnitude and pattern of these improvements varied. Overall, floating-point models experienced more substantial speedups than their quantized counterparts. However, the exact impact was more nuanced between the tested architectures. Across floating-point and 8-bit quantized RNNs (both with and without access to the oneDNN library), pruning reduced inference time and models followed a similar pattern. It should be noted that the quantized models were faster only than the floating-point models that did not use the oneDNN implementation. For the CNN architecture, the quantized models were faster only at the very beginning of the experiments. Moreover, the latency of the floating-point models decreased more steadily than that of the quantized ones. As the models became smaller, the more erratic graph of the quantized models’ latency diverged from the floating-point models’ graph even more, underlining the runtime overheads of the quantized kernels implementation. For the largest tested architecture, ResNet, during pruning, the quantized models remained faster than the floating-point models, up to about 10% of parameters left in the models. Beyond that point, a behaviour was observed that was similar to the one described for the CNN architecture.
Additionally, an interesting side finding emerged when we examined pruning patterns whose remaining structures aligned with multiples of eight. NNs with a dominant number of structures divisible by eight were faster than their slightly bigger or smaller versions, e.g. a layer with 32 output channels had a faster execution than a layer with 33 or 31 channels. This “hardware sweet spot” pointed to the phenomenon of a suboptimal configuration (Section
3.4) and the need to carefully consider the target hardware and runtime implementation while choosing the pruning pattern to obtain the lowest latency.
Surprisingly, pruning could improve the results obtained by the models, which was observed in the case of the RNN and the ResNet architectures. This behaviour could be attributed to the regularizing effect of pruning. Hence, it was not observed in the CNN, as it was regularized and quasi-optimally small, so further regularization only had adverse effects.
Finally, no significant differences were noted during the experiments between the
$L1$ and the
$L2$ methods that chose the structures to be pruned. These observations were similar to the results reported in Li
et al. (
2017). Instead, the number of pruning rounds, followed by fine-tuning, turned out to be a more important parameter. However, conclusions about the exact impact of pruning rounds remains an important direction for future work.
5.4 Knowledge Distillation
The knowledge distillation scheme utilized in the experiments had no direct impact on the model size or its inference latency. Instead, its primary benefit was improving the predictive quality of the base and pruned models (and therefore the quantized models), with only a few exceptions. In practical terms, this means that if a predefined quality acceptance threshold is present (e.g. a maximum allowable drop in accuracy), the use of knowledge distillation makes it possible to achieve more aggressive compression ratios (e.g. by applying pruning).
Our study deliberately focused on small networks of moderate depth. While response-based distillation, where the student mimics the output of the teacher, has proved to be effective in this context, prior work suggests that it may be less suitable for very deep NNs (Gou
et al.,
2021). In such cases, one might need to resort to other distillation schemes, e.g. matching activations from intermediate layers. Because the experiments did not include very deep models, a thorough investigation of alternative distillation paradigms remains beyond the scope of this work.
5.5 Limitations
Although this study covers various aspects of neural compression methods, its shortcomings must be realized. One significant limitation comes from the use of a single dataset, PTB-XL. It might introduce bias in the findings, which is associated with the specifics of the data, such as bias related to the demographics, the acquisition methods, or the representation of conditions. Another constraint arises from the runtime environment settings for the experiments. The inference latency results were obtained using a standard CPU, which may be too demanding for environments with strictly limited computing resources. Additionally, the experiments were performed using the full PyTorch runtime, which comes with requirements that may be unrealistic for tiny devices. For instance, it requires using an operating system or involves dynamic memory allocation. Other machine learning runtimes can be better suited for small computing devices, e.g. LiteRT (TensorFlow Lite) for Microcontrollers. While the shortcomings played a role in providing a clear and concise environment for experimentation, they might limit the representativeness of the results. However, the obtained results create a valuable proof of concept and motivation for further experimentation with the methods aimed at optimizing and deploying NNs on strictly resource-constrained devices.
6 Conclusion
In this study, three neural network compression techniques – quantization, structured pruning, and knowledge distillation – were applied to three different NN architectures used for ECG signal analysis. These NN compression methods were chosen because they address challenges associated with the use of NNs on devices with strict resource constraints, such as limited storage space or low computational capability.
The results indicated that model compression is a comprehensive problem in which the choice of techniques, software stack, and underlying hardware all play an important role. Quantization delivered smaller models, but the actual reduction was lower than the theoretical maximum. Its impact on inference speed was mixed, underscoring that low-bit representations alone are not always sufficient to guarantee practical efficiency gains. Pruning consistently reduced the model footprint and, in most cases, improved inference speed. However, it also revealed a strong dependence on hardware characteristics and pruning patterns. In fact, a poorly chosen pruning pattern could noticeably slow down a model. Knowledge distillation complemented both methods by improving the quality of the floating-point models and, consequently, the quality of their quantized versions.
Overall, neural network compression for resource-constrained devices is not a simple “plug-and-play” solution but rather a design space where effective strategies must be planned with awareness of the target runtime platform.
Finally, the compression techniques used in this study are not limited to ECG signal analysis. They are broadly applicable in other areas, such as autonomous systems, computational photography, or smart homes. However, their impact on model quality and resource consumption may vary in different domains due to factors such as the type of data, task complexity, inference requirements, or hardware architecture. Therefore, careful adaptation of the methods is crucial to ensure that they achieve their full potential when utilized in other application scenarios.